Hello readers! I’m eager to share some news with you: Perlara has signed new PerlQuests! One of our latest drug discovery efforts is a mucopolysaccharadosis (lysosomal storage diseases MPSIIIA and MPSIIIB) collaboration with Cure Sanfilippo Foundation (CureSFF). Sanfilippo syndrome is the common name for mucopolysaccharadosis. This post describes how the Sanfilippo syndrome PerlQuest came about, what it will entail, and what our next steps will be.
The path to PerlQuest
Over the past year, I’ve been working collaboratively with many families, foundations and academic institutions to explore drug discovery efforts for their diseases of interest. Together we discuss the model organism and drug screening possibilities, after which Perlara puts together an in-depth proposal, covering the details of the research plan. After the research plan is finalized, using the PerlArk™ platform, we begin with model organism genetic engineering, followed by model organism phenotyping and adapting to high-throughput screening. Once we identify screenable phenotypes, we embark on screening efforts to identify repurposed and novel compound rescuers of disease.
The Sanfilippo syndrome PerlQuest resulted from talks that began last February at the WORLD Symposium, when we reconnected with CureSFF to begin planning a PerlQuest launch for MPSIIIA and MPSIIIB.

What is Sanfilippo syndrome/MPSIII?
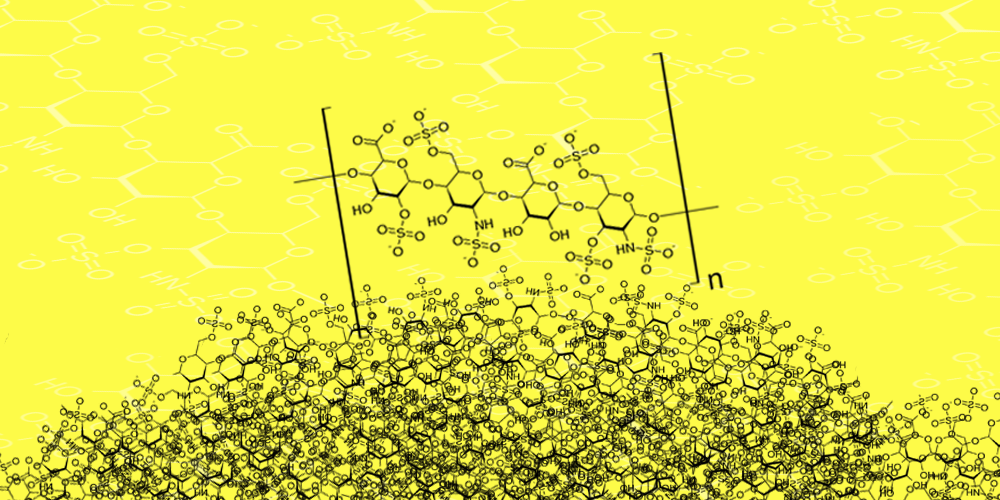
MPS stands for mucopolysaccharadosis, which is an accumulation of mucopolysaccharides (in this case, heparan sulfate) in lysosomes. MPSIIIA and MPSIIIB are caused by mutations in the SGSH and NAGLU genes, respectively. SGSH encodes sulfamidase (N-sulfoglucosamine sulfohydrolase) which is responsible for lysosomal degradation of heparan sulfate (a glycosaminoglycan or GAG – this will be important later!) and NAGLU encodes alpha-N-acetylglucosaminidase which is also responsible for the lysosomal breakdown of heparan sulfate. Mutations in both result in accumulation of heparan sulfate in the lysosomes, leading to neurological symptoms, aggressive behavior, dementia, seizures, hyperactivity and loss of vision.
Sanfilippo Syndrome PerlQuest: MPSIIIA
SGSH (MPSIIIA) is conserved in both nematodes and flies, with higher conservation in flies. In nematodes there are two orthologs, SUL-1 and SUL-3, and in flies the ortholog is the recently characterized CG14291 (Webber, 2018).
The nematode orthologs SUL-1 and SUL-3 are putative and uncharacterized. SUL-1 is 13% identical to the human protein and SUL-3 is 16.6% identical. We are eager to determine which SUL is appropriate to disrupt to model the human disease.
The fly protein, however, is 49.9% identical to the human protein sequence, which is highly homologous, and there is already published information on an MPSIIIA fly model. A paper was recently published on the MPSIIIA fly (Webber, 2018) demonstrating that RNAi knockdown of CG14291 (SGSH ortholog) leads to an accumulation of heparan sulfate and a disruption of lysosomal function and autophagy. Neuronal specific knockdown results in a progressive climbing defect.
Based on these data we are confident that the KO and pathogenic variants we are in the process of engineering will display phenotypes amenable to drug screening.

MPSIIIA fly climbing defect
Sanfilippo Syndrome PerlQuest: MPSIIIB
NAGLU (MPSIIIB) is highly conserved in both nematodes and flies, with similar protein sequence identity to the human protein. In nematodes, the ortholog is the uncharacterized K09E4.4, and in flies, the uncharacterized CG13397. The nematode ortholog is 31% identical to the human protein, and the fly ortholog is 37% identical to the human protein. Little is known about these proteins, but the high homology and phenotypes observed upon heparan sulfate accumulation in the MPSIIIA fly model provides confidence that a neurological phenotype will present itself in both of these models.
Our next steps
For both MPSIIIA and MPSIIB nematode and fly models, we just initiated organism engineering. We are eager to develop these models for use in Perlara’s drug discovery pipeline, and also provide as tools to the Sanfilippo research community.
Additionally, working on Sanfilippo alongside our PerlQuest for multiple sulfatase deficiency (MSD) – a collaboration with the MSD Action Foundation – will provide valuable information about the relationship between these two mucopolysaccharidoses, as MSD is related to MPSIIIA.
MSD is caused by mutations in SUMF1 (formylglycine-generating enzyme), which modifies sulfatases, activating them to breakdown sulfates. Sulfates are present on glycosaminoglycans, and when the GAGs are not broken down by the sulfatases such as SGSH, they accumulate in the lysosome. Pathogenic mutations in SUMF1 will lead to overall decreased sulfatase activities, including SGSH. It has also been shown that SUMF1 enhances sulfamidase activity when co-delivered to cells, and there is subsequent improvement of motor and cognitive function when administered to MPSIIIA mice.
We are thrilled to grow the Perlara family with the Sanfilippo syndrome PerlQuest, make more connections across disease indications which we call Organic Intelligence (OI), identify compounds for treatment of these diseases, and share data updates with the community!